Research
Over the past decade, technological advancements in next-generation sequencing and genomics have revolutionized the field of biology. Unfortunately, the unprecedented pace and volume of data being generated often exceeds the infrastructure and bioinformatic capabilities of many researchers. As a result, many disciplines have developed a divide between those actively involved in experimental research and data collection at the organismal level and those with the computational and bioinformatic skills to process these data. As a quantitative organismal biologist, my research continuously bridges this divide by integrating skills and training in computational genomics with experimental research in organismal biology while fostering collaborations with scientists on both sides. In particular, my research program utilizes these quantitative approaches to investigate the measurement, maintenance, and restoration of biodiversity (Fig. 1, below).
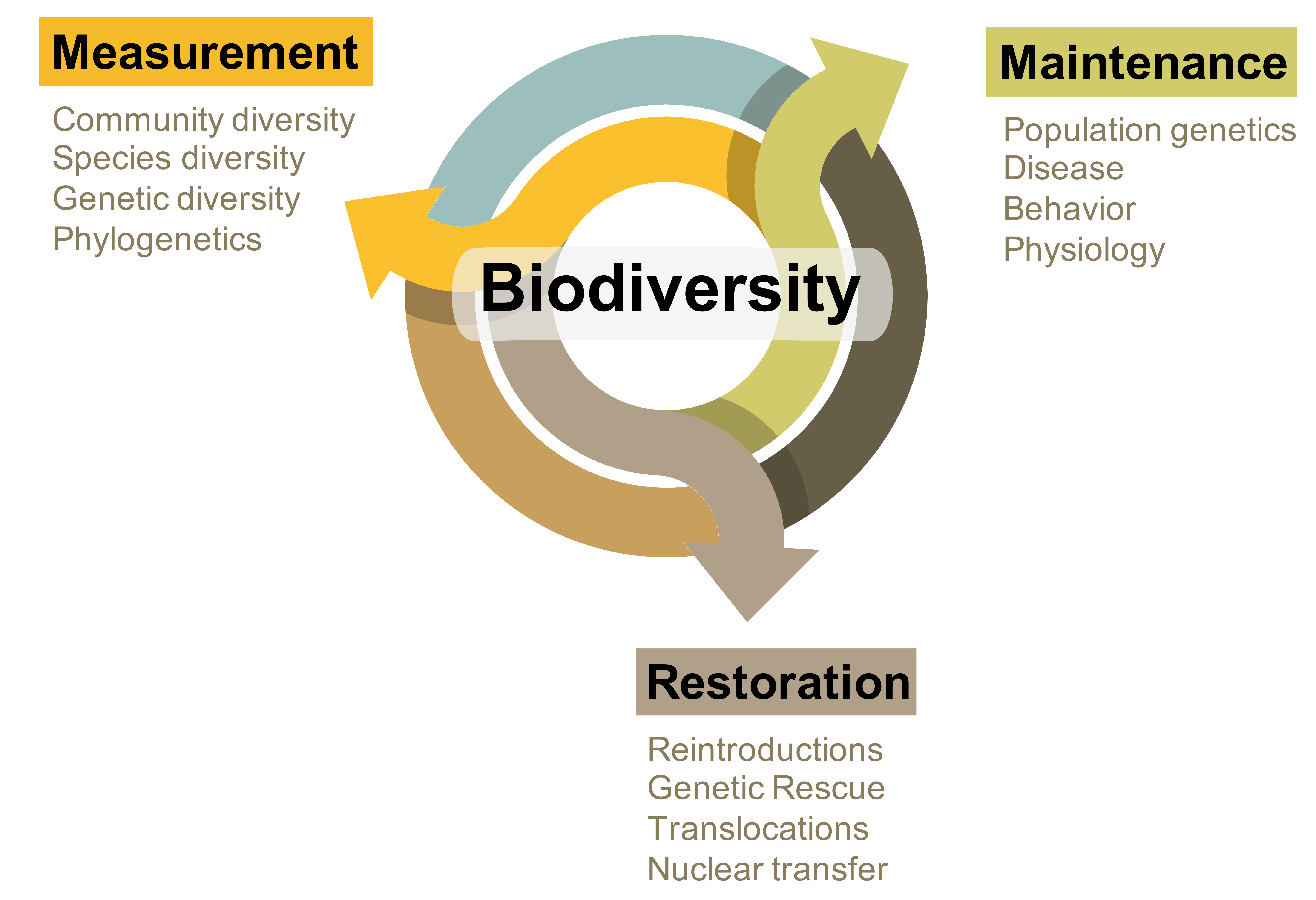
In one way or another, my lab’s various projects and interests intersect the chart in Fig. 1 (right), whether it be through the importance of understanding levels of genetic diversity, basic biology (e.g., physiology and behavior), or in directly translational results (e.g., population reintroductions). Furthermore, this paradigm is not limited to wild species, as many of the same concepts are directly applicable to domestic or otherwise agriculturally and economically important species. Below you will find specific examples of ongoing research projects in the lab.
Projects
-
Seagrass Strikes Back - Thermo-Priming for Marine Heatwave Resilience

Marine heatwaves (MHWs) are increasing in frequency, duration, and intensity, threatening the persistence of foundation species such as seagrasses. This project investigates whether thermo-priming, defined as controlled, sublethal heat exposure, can enhance thermal tolerance in shoal grass (Halodule wrightii). Using controlled mesocosm experiments, we exposed seagrass to simulated MHW conditions and quantified morphological performance (survival, growth, leaf area index) alongside transcriptomic responses using RNA-seq. A de novo reference transcriptome was assembled to identify differentially expressed genes associated with heat stress response. By integrating experimental ecology with functional genomics, this project aims to determine whether thermo-priming enhances resilience in seagrass and improves restoration success under predicted warming scenarios. Findings will inform climate-adaptive restoration strategies and contribute new molecular resources for an ecologically and economically critical coastal foundation species.
Funding: Seagrass Restoration Technology Development Initiative funded by the Mote Marine Laboratory and Florida Department of Environmental Protection
Collaborators: Linda Walters (UCF; co-PI), Carla Perscky (UCF; co-PI)
Assignees: Carla Perscky
-
Current and future impacts of a novel invasive parasite on Florida’s declining iconic pygmy rattlesnakes

This project investigates the impact of the invasive parasite Raillietiella orientalis (Ro) on Florida’s pygmy rattlesnakes, aiming to demonstrate that Ro is a primary cause of their recent population declines and to identify genetic factors associated with resistance to Ro and snake fungal disease (SFD). Emerging infectious diseases including Ro and SFD pose significant threats to North American snakes. Introduced by Burmese pythons, Ro is spreading rapidly in Florida, causing severe health issues and population declines in native snakes that already face SFD infections, particularly pygmy rattlesnakes. Project objectives include to (1) quantify Ro’s geographic spread and impact on pygmy rattlesnakes using data from ≥1,000 captures across 12 locations, (2) assess the relative importance of Ro, SFD, and co-infections on demographic declines, (3) conduct field surveys to compare genomic variation among individuals with different infection statuses, and (4) test whether MHC immunogenetic variation and/or genome-wide genetic variation predicts resistance and tolerance to Ro and SFD. Preliminary data include documented Ro infections and mortalities in pygmy rattlesnakes in central and south Florida but not in north Florida, identifying the current invasion front; observed clinical signs of SFD at all study locations; and genetic samples and morphometric data collected from 452 individuals. This study will confirm Ro prevalence using newly developed PCR assays effective on noninvasive fecal samples, detect SFD through skin swabs and lesion samples, perform whole-genome resequencing to identify genetic loci associated with disease resistance, and genotype MHC class II loci to link immune function with infection status. The results of this study will clarify the causes of pygmy rattlesnake declines, inform conservation efforts, and provide insights into the broader impacts of Ro on other snake species and potential zoonotic risks.
Funding: Morris Animal Foundation
Collaborators: Anna Savage (UCF; co-PI), Jenna Palmisano (UCF; co-PI)
-
The Ecological, Genetic, and Disease Impacts of Translocations on Florida Scrub-jays

This project will sample Florida scrub‐jays (Aphelocoma coerulescens, FSJs) throughout Brevard and Indian River counties (including surrounding the I‐95 Interchange to assess the ecological, genetic, and disease impacts of translocations. We will directly examine translocated birds including gathering demographic and disease data pre‐ and post‐translocation and testing whether genetic features of FSJs contribute to translocation success. The three objectives are to (1) translocate FSJs from sites of high bird density to sites with low density and available high‐quality habitat and monitor the survival, disease dynamics and integration success of these birds over a 2‐year period, (2) determine the genetic and immunogenetic compatibility of successfully translocated birds with the existing individuals, and (3) perform large‐scale molecular screening of pathogens to identify disease predictors and consequences of successful and unsuccessful reintroductions among the study sites. Collectively, we will use results from these objectives to better guide future FSJ management and translocation decisions throughout the eastern Florida corridor of FSJ habitat.
Funding: Florida Department of Transportation
Collaborators: Anna Savage (UCF; co-PI), Eric Hoffman (UCF; co-PI), David Breininger (UCF; co-PI)
Assignees: Gustavo Ortiz Vega
-
Geomagnetic Biogeography
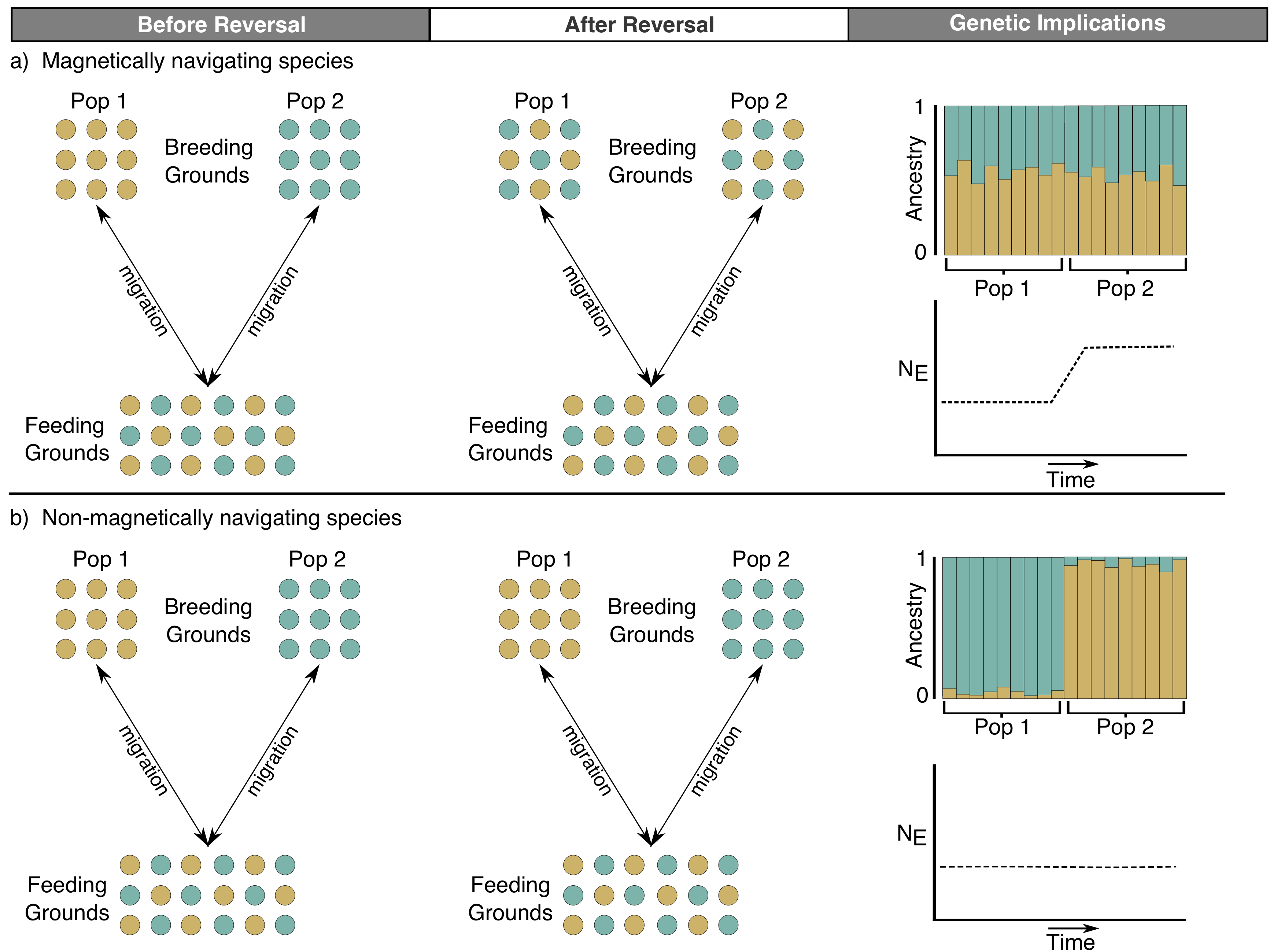
Earth’s magnetic field is quite dynamic - varying in strength, orientation, and even occasionally reversing polarity (swapping the North and South poles). It is possible this variation disrupts the navigation of species that depend on Earth’s magnetic field to migrate. Currently, we are outlining a conceptual approach for how population genetics can reveal changes in the demographic history and/or biogeography of species as a result of this variation - a phenomenon we call 'geomagnetic biogeography'.
We have provided some preliminary evidence in green sea turtles, where we recently showed that changes in gene flow between populations can account, at least partially, for a sudden increase in historical effective population size around the time of the last polarity reversal (780,000 years ago). This result, along with some great work by others, suggest that geomagnetic variation may have important consequences on the migration, and in turn, biogeography, of species that navigate using the geomagnetic field. Our goal is to continue work in this area in other 'magnetically sensitive' or navigating species to determine the role geomagnetic history plays in shaping species’ demography.Funding: N/A
Collaborators: Sönke Johnsen (Duke), Jay Wheeler (Duke), Roger Brothers (UNC), Ken Lohmann (UNC)
Assignees: TBD